Genome integrity is constantly threatened by DNA damage originating from endogenous and environmental sources. If not promptly repaired, DNA lesions can block DNA replication and lead to replication stress, genome instability and ultimately tumorigenesis. At the same time, cancer therapy involves DNA damaging therapeutics such as cisplatin and ionizing radiation that cause replication stress. As replication stress is toxic to cells with unchecked growth, it is exploited therapeutically to kill cancer cells. However, one of the main challenges in cancer therapy remains resistance acquisition. Our overreaching goal is to understand how cells deal with replication stress, and how it impacts genome and epigenome stability as well as sensitivity to environmental insults and cancer treatments.
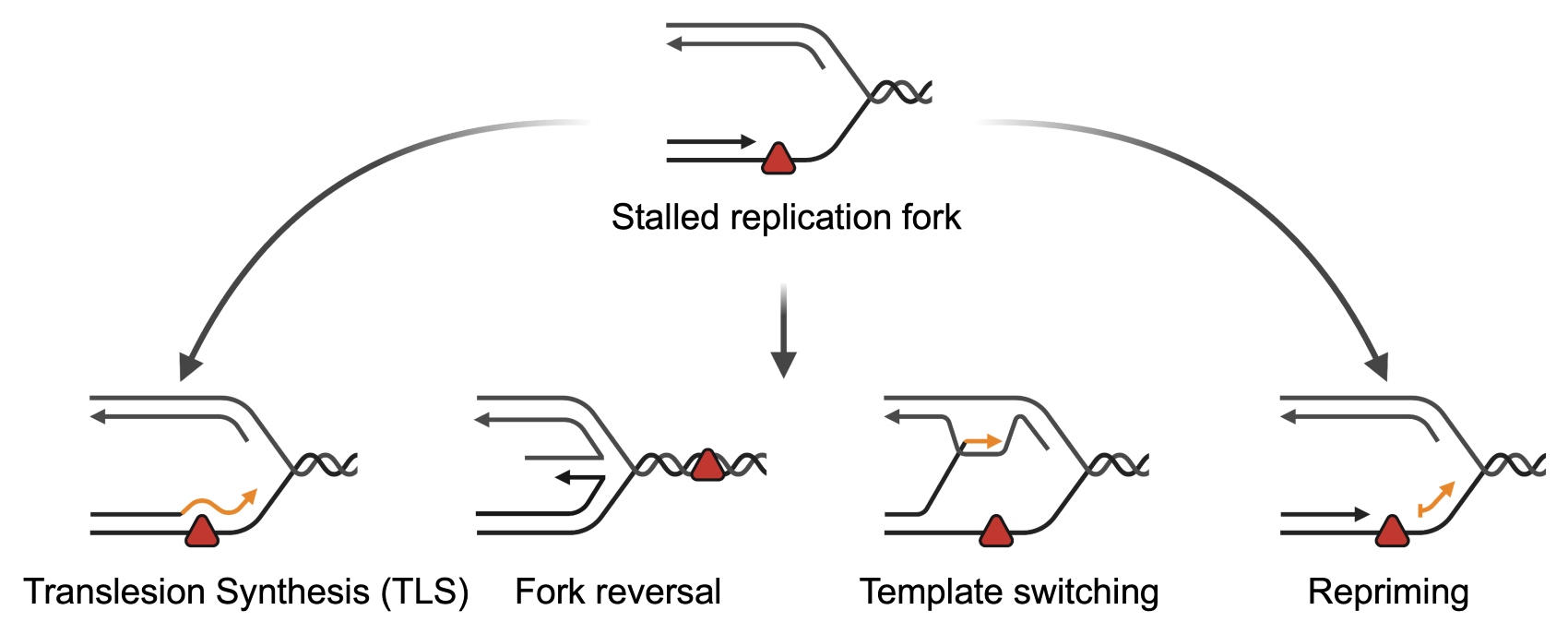
Cells have evolved mechanisms to respond to replication stress by tolerating DNA damage (Fig. 1). These mechanisms are a double-edge sword. On one hand, they ensure cell viability in conditions of replication stress. On the other hand, as they are potentially mutagenic, they can fuel carcinogenesis as well as contribute to the acquisition of resistance to DNA-damaging cancer therapeutics.
|
Fig. 1 Mechanisms of DNA damage tolerance (Adapted from Quinet et al., 2021, credit to Dr. Meroni)
|
Through the development of novel single-molecule approaches based on the DNA fiber assay (Fig. 2), we have uncovered that human cells frequently employ the Repriming mechanism to respond to replication stress, at the expense of accumulating post-replicative single-stranded DNA (ssDNA) gaps (Quinet et al., 2016, 2020). We have also started deciphering the molecular bases of gap-filling mechanisms in human cells and how they impact genome stability and cell sensitivity to environmental insults and cancer therapy (Quinet et al., 2016, Tirman, Quinet et al., 2021). 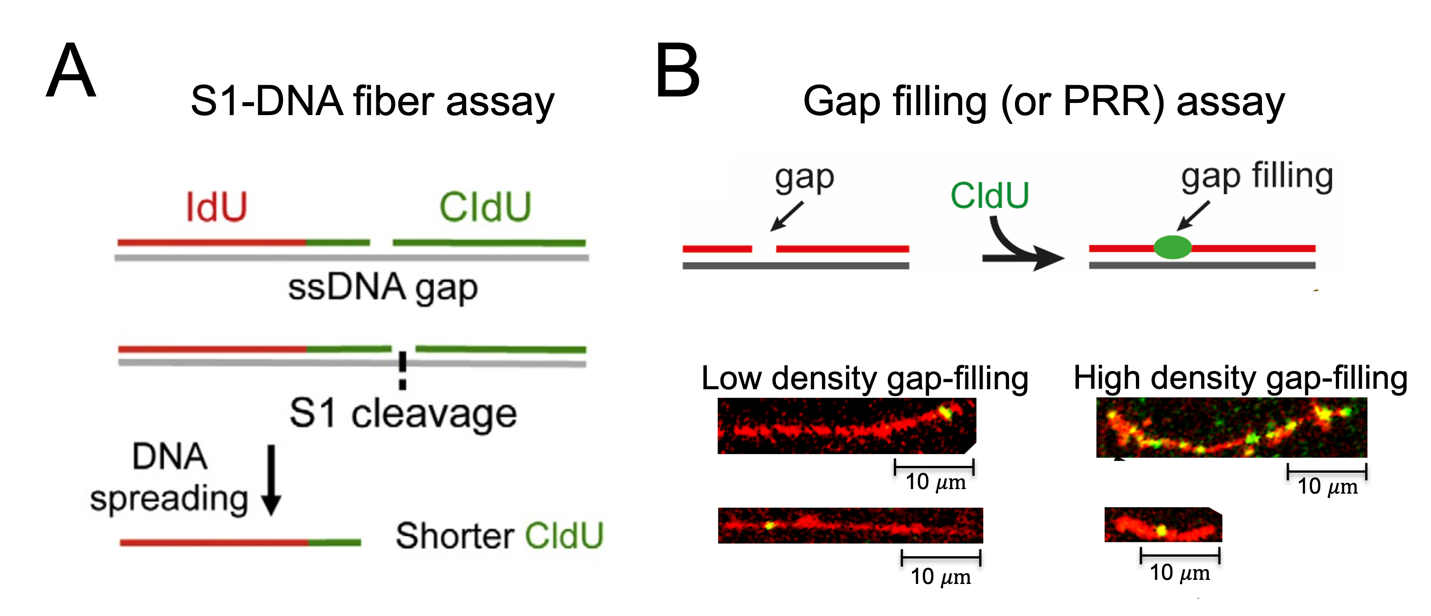
|
Fig. 2 Single-molecule microcopy approaches based on the DNA fiber assay to study post-replicative ssDNA gap formation and repair. (A) S1-modified DNA fiber assay. The use of the ssDNA-specific S1 nuclease on the DNA fiber assay enables detection of ssDNA gaps undetectable by standard DNA fiber protocols by generating shorter tracts that are used as a read-out for the presence of gaps. (B). Gap filling or Post-Replicative Repair (PRR) assay. A thymidine analog is added to the cell culture media to be incorporated during gap filling allowing direct visualization and quantification of these events. Images adapted from Martins, Tirman et al., 2022.
|
By combining our previously established approaches with cutting edge microscopy, proteomics, and next-generation sequencing using human cells, we now aim at:
- Defining the molecular steps of post-replicative ssDNA gap formation and repair
- Elucidating the crosstalk between post-replicative gaps and chromatin dynamics
- Investigating the response to replication stress induced by ionizing radiation and how it impacts cancer cells response to radiotherapy
Our long-term objective is to determine whether the expression of key replication stress response factors can be predictive of cancer cells response to therapy, and to identify novel strategies to improve current cancer treatments and overcome acquired resistance.
Our current projects are funded by an ATIP-Avenir grant (start date 01/01/2023), EDF (in collaboration with the team of Dr. Anna Campalans) and a Startup package for new Group leader from UMRE008/IRCM/IBFJ/CEA.