What types of screens can you perform?
Our platform has the technical capability to implement screens on human cell lines, preferably adherent, in miniaturized 96- and 384-well format. The cell lines used for siRNA and miRNA screenings must be transfectable with small oligonucleotides.
For cell-based measurements, we are equipped with a high content screening microscope allowing automated image acquisition and analysis in microplate format. This approach allows the phenotypic profiling of cells, i.e. the simultaneous quantification of multiple cellular features (cell morphology, fluorescence intensities, textures...) in 4 fluorescence channels + brightfield.
Our platform is also equipped with a multimode plate reader able to perform well-measurements in spectrophotometry, fluorescence, luminescence and dual-luminescence ("flash" and "glow"). Finally, thanks to a set of plate washers and fluidic dispensers, we also have the capacity to perform ELISA screenings.
We can support most common screening strategies:
- Direct screen, where effectors are directly assayed on the biological process of interest.
- Modifier screen, where the effectors are assayer with and without a modifying treatment (e.g. to assess the specific role of genes in the response to a radiotherapeutic treatment).
- Combinatorial screen, where effectors are assayed in pairs to explore their potential interactions. The STaRs Assay, which identifies the functional targets of a microRNA, is a form of combinatorial screening.
What is the STarS Assay?
Our platform has developed a screening approach allowing the identification of functional targets of a miRNA ("Suppressor Target Screen Assay" or "STarS assay", Fig. 3).
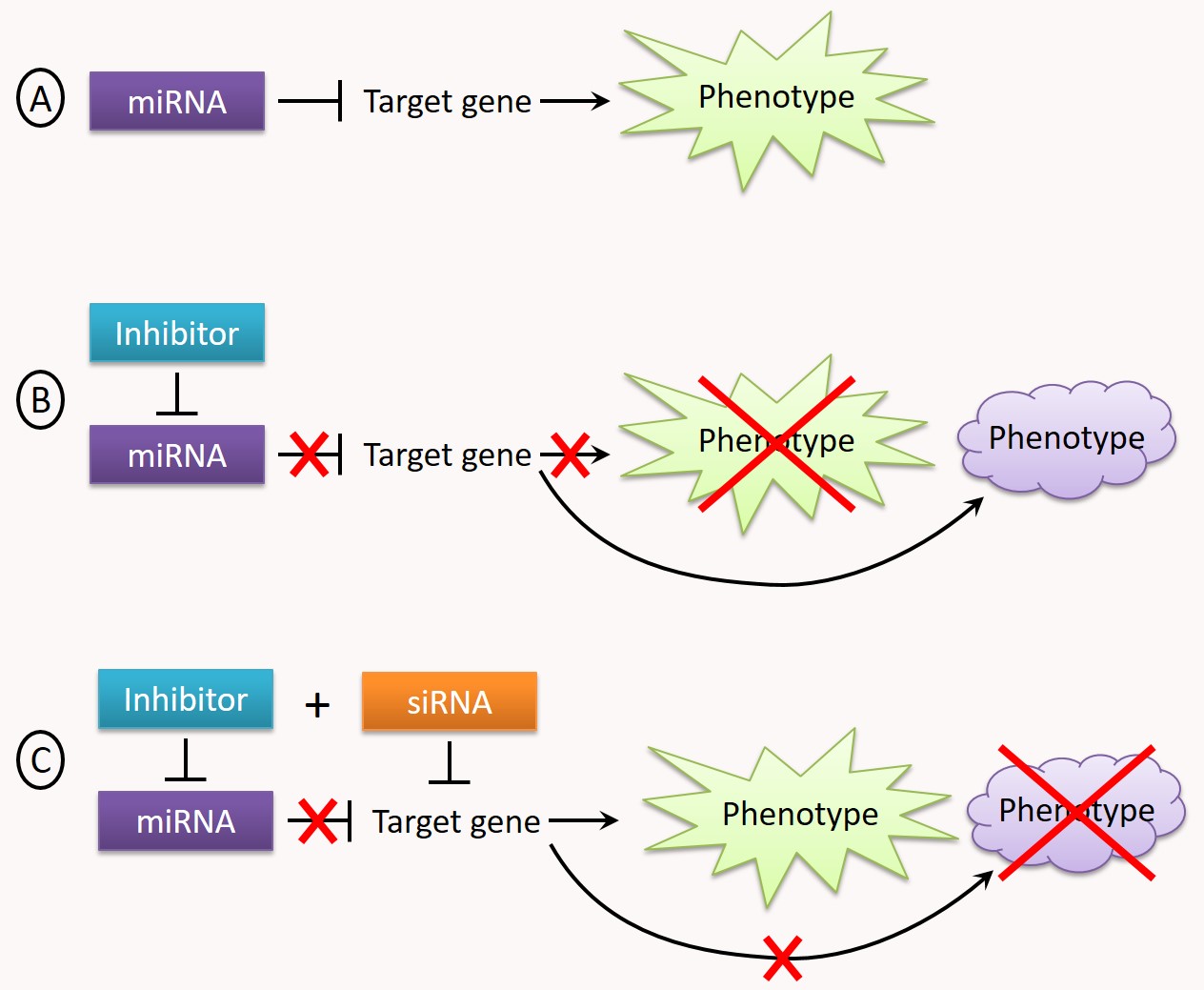
Figure 3: Identification of functional targets of a miRNA by "STarS assay". A. A miRNA regulates a cellular function by inhibiting the expression of one or more target genes, resulting in a "normal" cellular phenotype. B. When the miRNA is neutralized by an inhibitor (e.g. by a miRNA antisense), the expression of its target gene(s) is no longer repressed, resulting in a gain-of-function and a change in the measured phenotype. C. The STarS assay principle relies in screening a library of siRNAs, each of them individually paired with the miRNA inhibitor. At the end of such a screen, the set of genes targeted by the siRNAs capable of restoring - even partially - the initial phenotype constitute the functional targets of the miRNA.
What types of screens can't you do?
Concerning siRNA and miRNA screens, we have only human libraries, so we cannot accept, for the moment, screening projects on non-human cells. It is however possible to provide us with a library compatible with your cellular model, from which we can take over a screening project.
For sake of results reproducibility, we only work with cell lines. Any screening project on primary cells must be subject to a case-by-case feasibility study.
We cannot give an immediate positive answer to a project based on non-adherent cell lines, which are often difficult to transfect, and which limit the possibilities of imaging analysis. Such a project must also be studied on a case-by-case basis.
How does a typical screening project work?
Every screening project starts with a technical feasibility study, during which specifications are established and the overall cost of implementation is evaluated. Once all parties agree on the conditions for implementing the screen, the project can begin.
Experimentally, a screening project proceeds in several phases (Fig. 4)

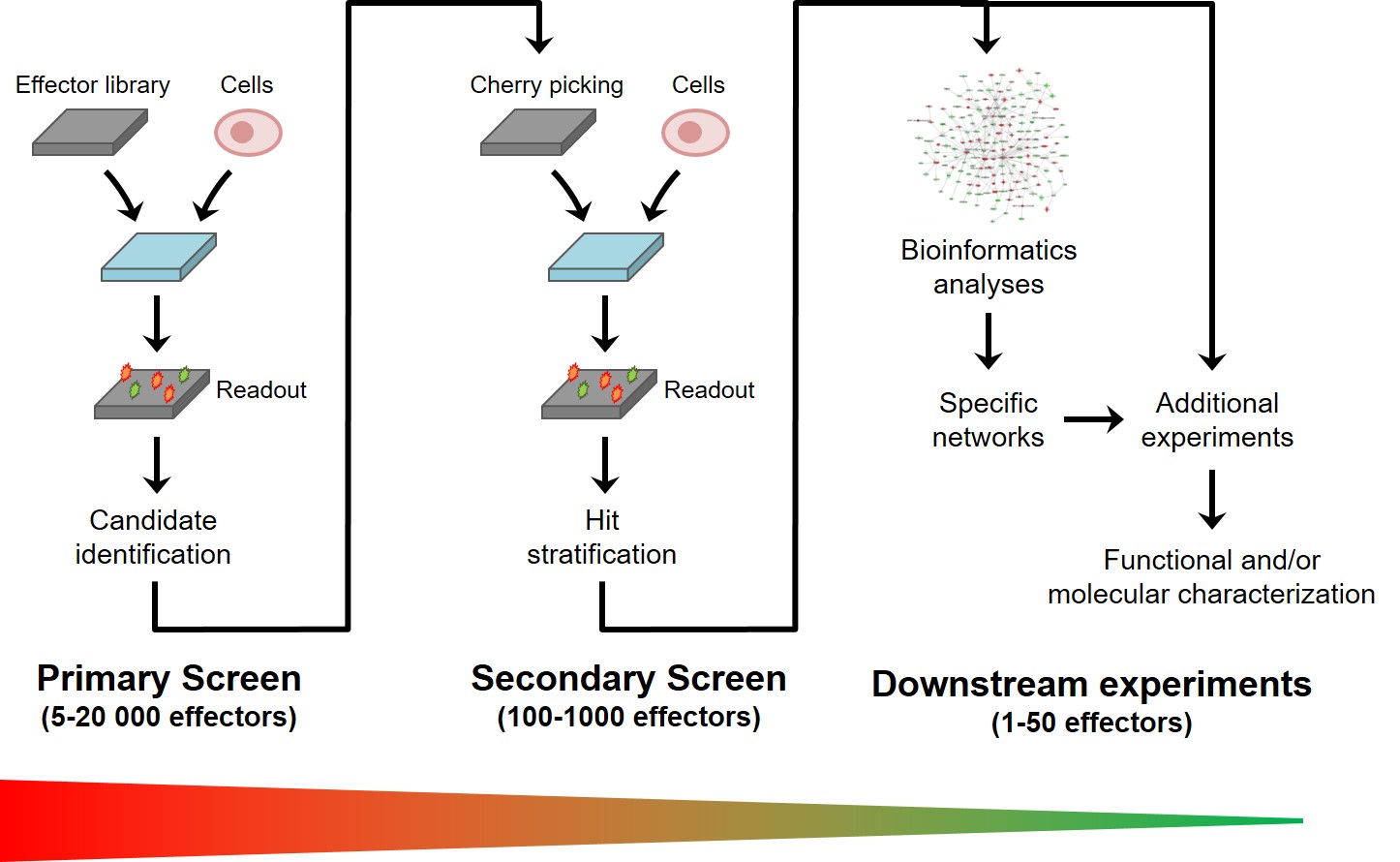
Figure 4: Structure of a typical screening project
The first step is to establish a miniaturized cell-based assay that is compatible with automation and sufficiently reproducible, specific and sensitive to ensure robust identification of candidate effectors. The duration of this step highly depends on the degree of maturity of the cell-based assay at the beginning of the project. It generally requires between 2 and 6 months of work, regardless of the nature or size of the library to be screened.
The screening itself always begins with a pilot screen (covering 5-10% of the library), during which the experimental conditions may be adjusted to improve the robustness of the screen. The duration of a screen depends mainly on the size of the library and the cell-based assay complexity. For example, a small screen (a few hundred of effectors) can usually be completed in a couple of weeks. However, a larger screen (such as a human genome-wide siRNA library screen) performed on a multiplexed cell-based assay may require up to 3 months of work.
Data processing and analysis allows for the identification of candidate effectors, which are then typically retested in a secondary screen, either on the same cell-based assay or on a complementary assay. Effectors that are confirmed to be active on the second assay constitute the "hit" list.
Depending on the library, additional analyses can also be performed based on the screening results (e.g., in the case of an siRNA screen, bioinformatics analyses can help identify specific signalling pathways from a hit list).
What is the typical price range for a screening project?
The cost of reagents and consumables for a human genome-wide siRNA screening project is usually comprised between 15 and 40 K€. A human genome-wide antisense miRNA or mimic screen (or a small drug library screen) usually requires between 2 and 10 K€ of consumables.
These costs are essentially dependent on the assay readout. For example, the cost of quantifying a constitutively expressed GFP is marginal; conversely, the quantification of a luciferase system can represent up to 70% of the screen costs.
Within the framework of an academic collaboration, only the costs of consumables and those related to the infrastructure (e.g. equipment maintenance) are invoiced.
Services for the private sector are subject to specific pricing that includes all operating costs (salaries, charges, equipment depreciation, environmental costs, etc.).
How soon can my project start?
The answer depends entirely on the degree of platform occupation at the time of the request. If the applicant laboratory has the funding, the average time to start a project is about 3 months after the first contact.
How many "hits" can I expect from my screening?
Experience shows that the final list of hits represents 1 to 3% of the screened library.