Our group is interested in systems and synthetic chemical biology. We develop in silico methods to search and design chemical and biological structures or networks constrained by experimental data. The applications of our work include reaction networks inference, structure activity/property relationships and rational molecular design.
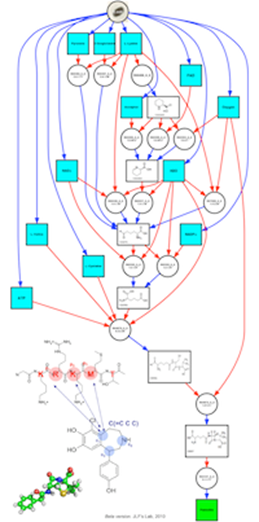
Our project within iSSB is to demonstrate that retrosynthesis, a widely used technique in chemistry, can be utilized to engineer chassis organisms such as E. coli or yeast to biosynthesize specific target compounds. Retrosynthesis analysis transforms a synthetic target product into precursors, following pathways to commercially available starting materials. The transformations are applied in the reverse direction from the actual synthesis. In the case of metabolic engineering, retrosynthesis consists of applying reversed biotransformations (i.e. reversed enzymes catalysed reactions) to a target product, following pathways to substrates that are endogenous to a chassis organism. In the case of biosensor engineering retrosynthesis is applied to effectors (small molecule binding to transcription factors) and pathways are enumerated between effectors and compounds to be detected such as disease biomarkers and pollutants. We are applying these techniques to engineer E. coli or yeast plasmids in order to construct combinatorial libraries of highest rank heterologous pathways found to produce target compounds. Combinatorial libraries are generated through the use of assembly PCR tested first in vitro, then in vivo. Production of the target metabolite is tested by measuring metabolites concentrations and fluxes.